Challenges in Imaging Expanded Samples
The resolution in light microscopy is limited by the diffraction limit of light, which means that structures that are closer than 200 nm apart cannot be distinguished unless researchers use Super-resolution techniques. An alternative approach to super-resolution methods was developed by Edward Boyden’s laboratory at MIT in 2015 (Chen et al., 2015). Instead of increasing the optical resolution of the microscope, Expansion Microscopy “expands” the sample isotopically. This technique has been gaining a lot of attention not only because it is an alternative approach to super-resolution, but also because it can be combined with current super-resolution techniques and increase even further the limits of what the microscopes “can see”. For a full summary of the protocols of expansion microscopy, please read Andor Technology technical article – What is Expansion Microscopy?.
Nevertheless, there are challenges to overcome to successfully image expanded samples. One of the main challenges is the loss of fluorescence induced by the different steps of the protocol. The complexity of the fixation procedure applied to the organelles causes problems in antibody labeling, leading to samples with a low signal. During the expansion procedure monomers of a polymer are delivered into the cells, and then, once inside, the polymerization is triggered (gelation). During gelation, the inefficient crosslinking results in fragmented signal preservation in the hydrogel. Next, the homogenization step aims to avoid sample distortion and ensure that the specimen organization is kept intact. However, the fluorescence signal can decrease up to 60% during homogenization. During the actual expansion, a 4-fold expansion means a 64-fold volumetric expansion; hence the fluorophore density and brightness of the fluorescence signal is lowered by 64x. Therefore, imaging expanded samples requires an instrument that is sensitive enough to detect low light signals. Other challenges consist of dealing with the expansion itself including imaging a very large field of view (Figure 1.), and imaging deep into the sample. These requirements are problematic and difficult to achieve with conventional microscopes.
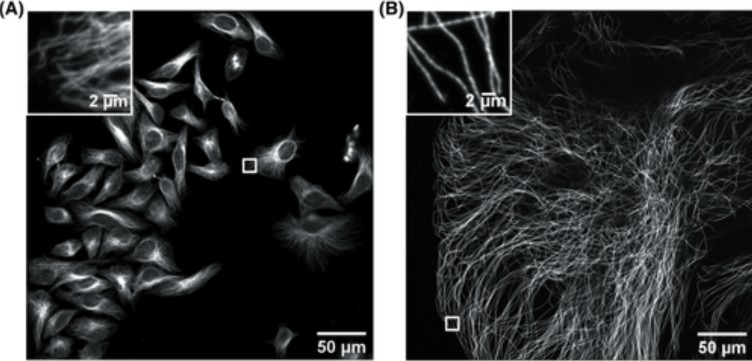
Figure 1. Confocal image of HeLa cells non expanded microtubules (right) and 4.5x linearly expanded microtubules (left). Imaging was performed on an Andor high-speed spinning-disk confocal microscope (Dragonfly) with a 40×, numerical aperture (NA) 1.15 water- immersion objective. (A) Confocal image of HeLa cells with immunostained microtubules imaged at a single xy plane at the bottom of the cells. The inset in the upper left zooms in on the small box at the middle right. (B) Confocal image of a ∼4.5× linearly expanded HeLa cell with immunostained microtubules imaged at a single xy plane at the bottom of the cell. The inset in the upper left zooms in on the small box at the bottom left. Scale bars in (B) indicate post-expansion scales. Only a fraction of an expanded cell fills the entire field of view. The respective insets display a zoom of the respective small boxes of the full field of view. (Zhang, C., et al, Current Protocols in Neuroscience,2020).
Technology Solutions for Imaging Expanded Samples
To overcome the loss of signal in expanded samples, a highly sensitive acquisition system is needed. Researchers need to use a microscope that can facilitate optimal delivery of light to the samples and extremely efficient capture of all the photons from the sample, i.e. high QE detectors. The use of highly sensitive cameras turns camera-based confocal systems into the ideal solution to capture all signals from expanded samples, even the dimmest ones.
In addition, to image expanded samples with high productivity, a large field of view of acquisition is a must-have. A multipoint confocal microscope that delivers images in an instant with a large FOV of acquisition is, therefore, the perfect solution. The multipoint confocal will need to have optimized pinhole spacing to avoid pinhole-crosstalk, allowing imaging deep into thick expanded samples. Moreover, thick expanded samples are big, and on many occasions, multiple tiles are required to be able to gather all the three-dimensional information on the sample like the example displayed in figure 2. Ideally, the system should have a very flat uniform illumination that when used in combination with software, can deliver seamless stitching.
e use of multiple wavelengths is an important requirement that will allow users to label the numerous targets in the sample and obtain information on the localization of different structures and how they relate to each other. The use of Near Infra-Red (NIR) wavelengths is an extra bonus, and not only extends the fluorochrome pallet but, very importantly, NIR wavelengths penetrate deeper into the sample.
Additionally, imaging the expanded sample using super-resolution techniques will afford improved resolution. For example, combining Expansion Microscopy with SRRF (super-resolution radial fluctuations) will give an increase of at least an extra 2X on the resolution obtained with the expanded sample alone.
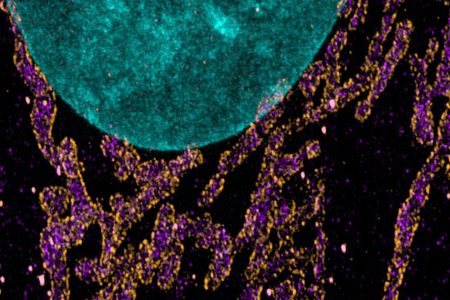
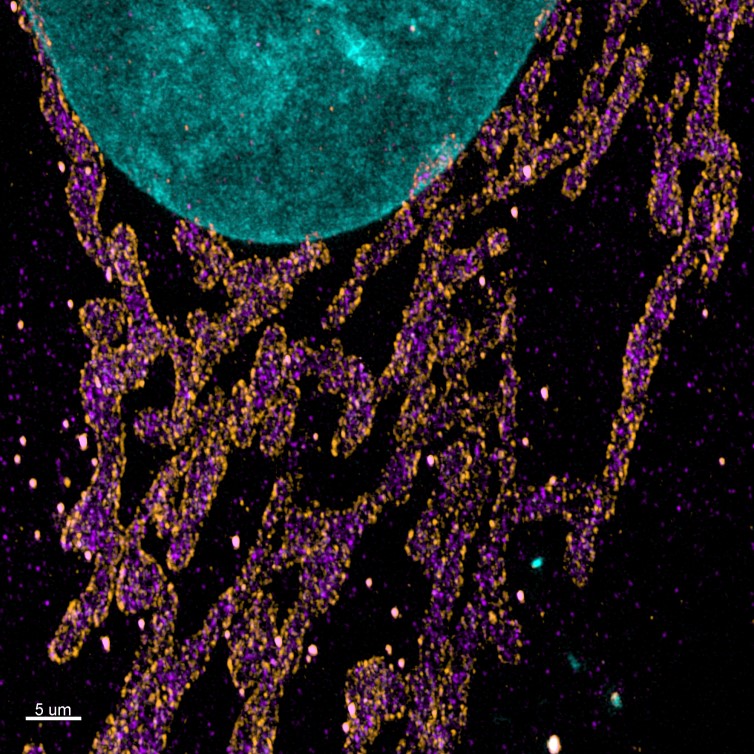
Figure 2. Maximum projection images of the mitochondrial import receptor. Cell where labelled using the post-expansion protocol with HSP70 (cytosolic side of organelles membrane – purple), TOM20 (outer mitochondrial membrane – orange), DAPI (nucleus – Cyan), and image with Andor Dragonfly. Courtesy of Line Verckist,University of Antwerp.
Andor Solutions for Image Expanded Samples
Andor recommends the Dragonfly confocal equipped with either the iXon Ultra 888 back-illuminated EMCCD, or the Zyla 4.2P sCMOS cameras for expansion microscopy. The dual lens spinning disk system, combined with highly sensitive EMCCDs and sCMOS detectors delivers a high-quality image even in samples with weak staining. Dragonfly has optimised pinhole spacing which allows imaging deep into samples without pinhole crosstalk. Further, Andor’s patented Borealis illumination delivers uniform illumination across the field of view, which is a key factor for imaging the multiple tiles on a large expanded image. Combined with a large field of view,22 mm diagonal, Dragonfly can deliver fantastic images with remarkable productivity.
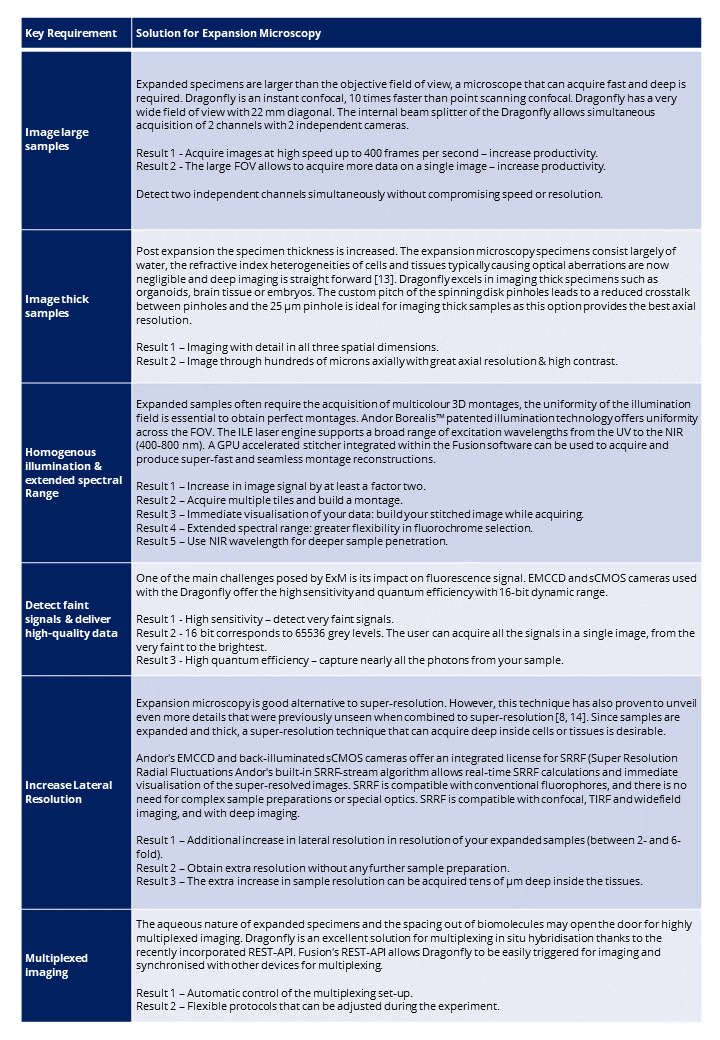
In the following table, we present how dragonfly spinning disk confocal addresses the current and future challenges in imaging expanded samples
References:
- Chen, F., P.W. Tillberg, and E.S. Boyden, Expansion microscopy. Science, 2015.
- Zhang, C., Kang, J. S., Asano, S. M., Gao, R., & Boyden, E. S. Expansion microscopy for beginners: Visualizing microtubules in expanded cultured HeLa cells. Current Protocols in Neuroscienc,
Original article: https://andor.oxinst.com/learning/view/article/dragonfly-the-ideal-confocal-system-for-expansion-microscopy